Logistic regression tutorial¶
This part of tutorial is derived from its step-by-step notebook version multinomial logistic regression example, the emphasis is to showcase the basic capacity of MinPy.
We will work on a classification problem of a synthetic data set. Each point is a high-dimentional data in
one of the five clusters. We will build a one-layer multinomial logistic regression model. The goal is to learn
a weight matrix weight, such that for a data point x the probability that it is assigned to its class (cluster)
is the largest.
The data is generated with the following code.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 | import numpy as np
""" Generates several clusters of Gaussian points """
def gaussian_cluster_generator(num_samples=10000, num_features=500, num_classes=5):
mu = np.random.rand(num_classes, num_features)
sigma = np.ones((num_classes, num_features)) * 0.1
num_cls_samples = num_samples / num_classes
x = np.zeros((num_samples, num_features))
y = np.zeros((num_samples, num_classes))
for i in range(num_classes):
cls_samples = np.random.normal(mu[i,:], sigma[i,:], (num_cls_samples, num_features))
x[i*num_cls_samples:(i+1)*num_cls_samples] = cls_samples
y[i*num_cls_samples:(i+1)*num_cls_samples,i] = 1
return x, y
|
The visualization of the data:
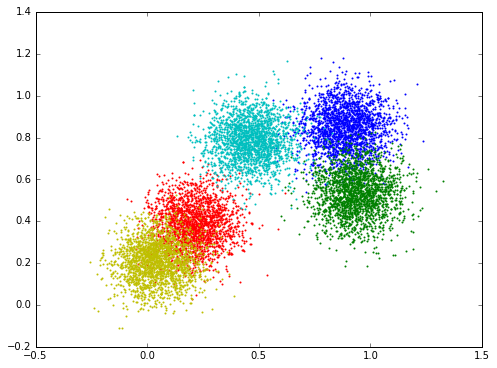
The following is the numpy version. The function predict outputs the probability, the train function iterates
over the data, computes the loss, the gradients, and updates the parameters w with a fixed learning rate.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 | import numpy as np
import numpy.random as random
from examples.utils.data_utils import gaussian_cluster_generator as make_data
# Predict the class using multinomial logistic regression (softmax regression).
def predict(w, x):
a = np.exp(np.dot(x, w))
a_sum = np.sum(a, axis=1, keepdims=True)
prob = a / a_sum
return prob
# Using gradient descent to fit the correct classes.
def train(w, x, loops):
for i in range(loops):
prob = predict(w, x)
loss = -np.sum(label * np.log(prob)) / num_samples
if i % 10 == 0:
print('Iter {}, training loss {}'.format(i, loss))
# gradient descent
dy = prob - label
dw = np.dot(data.T, dy) / num_samples
# update parameters; fixed learning rate of 0.1
w -= 0.1 * dw
# Initialize training data.
num_samples = 10000
num_features = 500
num_classes = 5
data, label = make_data(num_samples, num_features, num_classes)
# Initialize training weight and train
weight = random.randn(num_features, num_classes)
train(weight, data, 100)
|
The minpy version is very similar, except a few lines that are highlighted:
- Among some new imported libraries,
minpy.numpyreplacesnumpy. This lightweight library is fully numpy compatible, but it allows us to add small instrumentations in the style of autograd - Defines loss explicitly with the function
train_loss - MinPy then derives a function to compute gradients automatically (line 24)
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 | import minpy.numpy as np
import minpy.numpy.random as random
from minpy.core import grad_and_loss
from examples.utils.data_utils import gaussian_cluster_generator as make_data
from minpy.context import set_context, gpu
# Please uncomment following if you have GPU-enabled MXNet installed.
# This single line of code will run MXNet operations on GPU 0.
# set_context(gpu(0)) # set the global context as gpu(0)
# Predict the class using multinomial logistic regression (softmax regression).
def predict(w, x):
a = np.exp(np.dot(x, w))
a_sum = np.sum(a, axis=1, keepdims=True)
prob = a / a_sum
return prob
def train_loss(w, x):
prob = predict(w, x)
loss = -np.sum(label * np.log(prob)) / num_samples
return loss
"""Use Minpy's auto-grad to derive a gradient function off loss"""
grad_function = grad_and_loss(train_loss)
# Using gradient descent to fit the correct classes.
def train(w, x, loops):
for i in range(loops):
dw, loss = grad_function(w, x)
if i % 10 == 0:
print('Iter {}, training loss {}'.format(i, loss))
# gradient descent
w -= 0.1 * dw
# Initialize training data.
num_samples = 10000
num_features = 500
num_classes = 5
data, label = make_data(num_samples, num_features, num_classes)
# Initialize training weight and train
weight = random.randn(num_features, num_classes)
train(weight, data, 100)
|
Now, if you uncomment line 9 to set MXNet context on GPU 0, this one line change (set_context(gpu(0))) will enable the same code to run on GPU!
For more functionality of MinPy/MXNet, we invite you to read later sections of this tutorial.